Watching it grow: developing a digital embryo Understand article
What if you could witness the development of a new life, taking your time to study every detail, every single cell, from every angle, moment by moment? Sonia Furtado talks to the scientists who made this possible by creating a digital zebrafish embryo.

Jochen Wittbrodt and Annette Schmidt
Image courtesy of EMBL Photolab
Imagine Google Earth™ took new satellite photos at regular intervals, so that as well as viewing the whole planet and zooming in on different countries or cities, you could play back the sequence of photos and see what had changed over 10 or 20 years. Replace Earth with a developing zebrafish embryo, collect a few hundred thousand snapshots recorded over 24 hours, and you get what scientists at the European Molecular Biology Laboratory (EMBL)w1 have dubbed ‘the digital embryo’.
Biophysicist Philipp Keller, biologists Annette Schmidt and Jochen Wittbrodt, and physicist Ernst Stelzer teamed up to develop a technique for obtaining a 3D representation of the first 24 hours of a zebrafish embryo’s development (Keller et al., 2008). It all started in early 2006, when a fellow scientist suggested that Philipp simply put an embryo in the microscope and track all the cells. Prompted by this conversation, Philipp took a fresh look at single-plane illumination microscopy (SPIM) (see glossary for all italicised terms), a 3D imaging technique previously developed by Ernst’s group. He refined the basic principles of SPIM to record embryonic development; the result was digital scanned laser light sheet fluorescent microscopy (DSLM).
Because cells in a developing embryo are constantly dividing and moving, snapshots must be taken at very short intervals to track the changes. This requires a microscope which combines high imaging speed with high image quality, to distinguish between cells sitting very close to each other. And, of course, in order to follow the embryo’s development, the cells must be kept alive and behaving normally over a period of one or two days, and the microscope must not damage the fluorescent stain used to mark the cells.
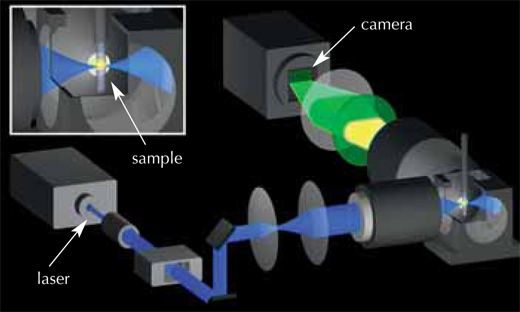
Image courtesy of Petra Riedinger
The solution was to use a very thin laser beam, minimising the damage to both the stain and the embryo. This beam shines through the embryo, exciting the fluorescent stain in the cells and making it emit light. The laser beam moves in a vertical line, and the fluorescence emitted from the sample at a 90° angle to the laser beam is detected by a camera, creating an image slice thousandths of a millimetre thin (see movie 1w2). The embryo is then moved a tiny horizontal step, and the next image slice is scanned. Once the whole embryo has been scanned from one viewpoint, it is rotated by 180° and scanned again, so that a full 3D image can later be composed. The embryo lives and carries on developing, which means the ‘slices’ can be repeated periodically on the same embryo and compared to track the changes. The EMBL scientists ‘sliced’ their zebrafish embryos either every 60 or 90 seconds over a 24-hour period, obtaining around 400 000 images per embryo.
The next challenge was to devise a way to analyse all three terabytes of data for each embryo. The scientists chose an automated approach: they used clusters of computers working in parallel, both at EMBL Heidelberg and at the Karlsruhe Institute of Technology, Germanyw3. “Each computer is given one snapshot of the embryo, and told to look for cellular nuclei in that image,” Philipp explains. Each nucleus represents a cell, so by combining the information for all slices, scientists can generate their digital embryo: a visual representation of all the embryo’s cells, where they are at a given point in time, where they move to next, and when and where they divide. The result is a 3D time-lapse video of the developing embryo.
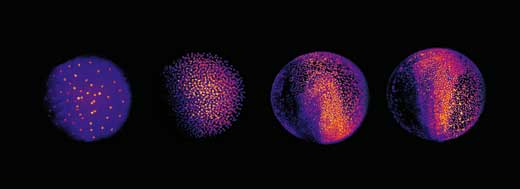
Image courtesy of EMBL Photolab
This enabled Philipp, Annette and Jochen to shed new light on different stages of the embryonic development of zebrafish. At a very early stage, the zebrafish embryo is basically a group of cells sitting on top of the yolk sac (the embryo’s nutrient source). At first, the cells divide in a wave that moves out from the centre in all directions, like a ripple in a pool. The reconstruction showed that after the first nine cell cycles, the pattern of cell division changes to a wave that moves out from the centre in only one direction and then splits up to continue along the periphery in two half-circles (see image below and movie 10w2). The line between these two half-circles will later become the animal’s body axis, defining the body’s symmetry. So the scientists discovered that the zebrafish’s body axis is defined at an earlier stage in its development than was previously supposed, at a time when maternal genes (in the form of mRNA deposited in the egg) are still the sole blueprint used for protein production.
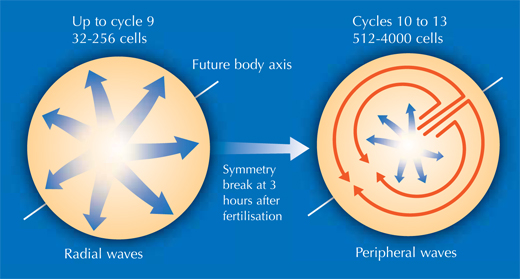
Image courtesy of Nicola Graf
The researchers also studied gastrulation, the process by which cells from the embryo’s original single outer layer migrate inward to form the remaining two germ layers, all three of which will eventually give rise to different types of tissue. “It takes textbooks two or three pages to describe this process, but when you see the movies of the digital embryo, it’s suddenly quite easy to understand,” Annette says. And what’s more – she and her colleagues discovered that the textbooks are actually wrong.
There has been an ongoing, sometimes fervent, debate in the field since the 1980s over how cells migrate to form these layers. The prevailing textbook view was that cells involute – that is, they migrate to an opening in the embryo and roll over the margin to form a layer underneath, like the edge of a swimming cap rolling up around your head, on the inside. Yet some scientists argued that individual cells on the outer layer simply dive in, or ingress, right where they are, to form a layer beneath.
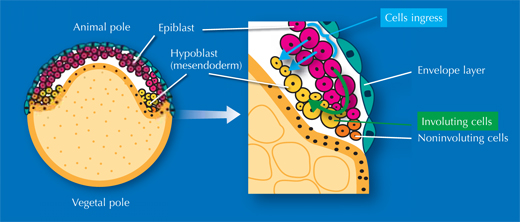
Image courtesy of Nicola Graf
”We really settled the issue nicely,” says Jochen. “It turns out everyone was right!” It’s a question of where you look. On one side of the embryo, cells involute, whereas on the other side, they ingress (see movie 16w2). The controversy had arisen because different scientists were looking at different areas of the embryo. Since they couldn’t watch the whole process unfold, they were, however, unable to determine exactly which area they were looking at. In overcoming these constraints, the EMBL scientists were able to uncover the truth.
The researchers have also used the digital embryo as a developmental blueprint, to find out where the cells contributing to a specific organ or tissue come from. As a first example, they used the eye. In the digital representation, at an advanced developmental stage (i.e. late in the 24-hour video), the scientists marked the cells that they knew were involved in forming one of the zebrafish’s eyes. They then tracked the cells back in time to find out where they originated (see movie 11w2).
The scientists have made their digital embryos publicly available on the Internetw2, along with tools for other scientists to analyse their own microscopy data. And what next? With Google Earth, as well as seeing the whole planet and zooming in on different places, users can add their own notes and markers and view those added by other people. Similarly, Philipp, Annette, Jochen, and Ernst envision their digital embryos becoming what they call ‘virtual embryos’: resources in which other scientists can view developmental processes, zoom in for more information, and add their own annotations and results. In the long run, the scientists would like to expand the digital embryo’s scope to other species, as this would enable scientists to quantitatively compare how different embryos develop, which would provide valuable insights into evolution. The digital embryo has much room for growth, and with growing numbers of people willing to help it develop, it appears to have a promising life ahead.
Glossary
Digital scanned laser light sheet fluorescent microscopy (DSLM): An improved version of SPIM, which uses a thin laser beam rather than a full light sheet, thus reducing damage to both specimen and fluorescent dye.
Gastrulation: The phase in early embryonic development during which the three germ layers are formed: ectoderm, mesoderm and endoderm. The timing and molecular mechanism of gastrulation differ between organisms.
Genetic strain: A genetically uniform group of animals, used in laboratory experiments. A genetic strain can be developed by inbreeding, mutation or genetic engineering.
Single-plane illumination microscopy (SPIM): This method allows 3D observation of processes in living organisms, even in deep tissue layers. It detects fluorescence at an angle of 90° relative to the axis of illumination with a sheet of laser light, permitting optical cutting. The specimen is not positioned on a microscope slide but in a liquid-filled chamber which is rotated during observation.
The zebrafish as a model organism
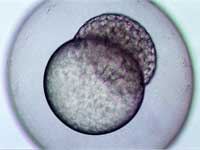
stage, two hours and forty-five
minutes after fertilisation
Image courtesy of Angharad Jones
, Wellcome Images
For ethical reasons, we can’t easily study early (embryonic) development in humans. Conveniently though, the process is similar in all vertebrates, so other animals are used as model organisms to understand our own development. But why zebrafish? We owe it all to George Streisinger, a Hungarian biologist at the University of Oregon, USA. At the beginning of the 1980s, he dreamed of applying the molecular techniques he had previously used for the study of viruses to investigate the genetics and development of a vertebrate. As a fish hobbyist, he chose to work on zebrafish, which he knew were easy to keep and breed. There are a number of practical reasons which make zebrafish an attractive model system: firstly, they are small enough to keep the large numbers required for genetic studies, yet big enough to do classical embryological manipulations such as transplantations.
They have a relatively short generation time (3-4 months), produce large numbers of embryos (100-200 per mating) and provide easy access to all developmental stages due to external fertilisation. Because the embryos are optically transparent and develop rapidly, with all the important structures of an adult fish being established after three days, you can easily study their development – using either a simple light microscope or more complex technology, as described in this article. In addition, the zebrafish genome has recently been sequenced, and defined genetic strains are available, offering a unique opportunity to study not only embryology, but also genetically inherited diseases and the genetics underlying developmental biology – Streisinger’s dream has come true.
References
- Keller PJ, Schmidt AD, Wittbrodt J, Stelzer EH (2008) Reconstruction of zebrafish early embryonic development by scanned light sheet microscopy. Science 322: 1065-9. doi: 10.1126/science.1162493
Web References
- w1 – To learn more about EMBL, the European Molecular Biology Laboratory, see: www.embl.org
- w2 – The Digital Embryo website includes videos showing the development of the zebrafish embryo, as well as data to download: www.digital-embryo.org
- Movie 1 is a schematic representation of the DSLM’s operation principle.
- Movies 2 and 3 show the development of a zebrafish embryo during the first 24 hours.
- Movie 10 illustrates the symmetry break in the cell divisions after nine cell cycles, and the early determination of the embryo’s body axis.
- Movie 11 shows the migratory tracks of cells forming the early zebrafish eye.
- Movie 16 illustrates how involution and ingression both play a role in zebrafish gastrulation.
- w3 – For more information about the Karlsruhe Institute of Technology, Germany, see: www.kit.edu
Resources
- Patterson L (2010) Getting ahead in evolution. Science in School 14: 16-20.
- Hodge R (2006) A search for the origins of the brain. Science in School 2: 68-71.
- For a collection of learning and research resources on embryonic development, see: http://people.ucalgary.ca/~browder/virtualembryo
Institutions
Review
This article describes cutting-edge advances in developmental biology. The potential outcome –a virtual embryo which could be shared amongst and annotated by other scientists, just as photos and comments are added to Google Earth – is an exciting example of how research findings can be disseminated, commented upon and, perhaps, extended. This could link well with the ideas about global science and communication that appear in some syllabuses.
There is much in this article that could enthuse interested biology students. The use of sophisticated microscopy in pushing forward scientific boundaries could give rise to discussions about how technology is holding back discoveries, or about how much scientific ‘knowledge’ is ephemeral and uncertain, being dependent on current technology.
The use of zebrafish embryos could also give rise to an ethical debate. Is it right to experiment on fish embryos? What about mammalian embryos? What about human embryos? This could lead to a discussion of some of the legal aspects of this kind of research.
Generally, the article could be useful for biology teachers to update their own knowledge. It may also be used as background reading – possibly for students entering biology olympiads or those with interviews for university places. In addition, the video footage is helpful to illustrate embryonic development for any courses that include this topic.
Suitable comprehension questions include:
- Describe and discuss any ethical issues involved in the work carried out by the scientists.
- Suggest how the work on fish embryos could be of relevance to treating human diseases in the future.
Sue Howarth, UK





