From model organism to medical advances Understand article
A simple fungus used to brew beer is now used around the world to advance cancer research.

Cancer affects millions of people worldwide each year. However, thanks to ongoing research into cancer and the underlying cell biology, treatments and survival rates are improving all the time. Perhaps surprisingly, the yeast Schizosaccharomyces pombe is an important tool for cancer researchers. This well-studied model organism (see box) has enabled groundbreaking, Nobel Prize-winning discoveries and for more than 50 years has provided insights into how both normal and cancerous cells grow and divide.
Cancer is the human equivalent of a weed – unwanted abnormal cells grow out of control and in the wrong place. It is often the result of a series of changes in the DNA of a cell, known as mutations, that accumulate over time. These mutations can make cells grow too fast and divide too frequently, eventually forming tumours that invade nearby tissues and organs and cause damage. To understand how these mutations cause cancer, it is first important to understand how the cells work under normal circumstances.
Model Organisms
Many basic life processes, particularly at the cellular level, are almost identical in all living things. This means that scientists can use organisms such as fruit flies, zebrafish, mice or even yeast to study the fundamentals of cell growth and development and to apply that knowledge to humans. Certain species have proven particularly useful for research, so they have become known as model organisms.
These organisms tend to be small, robust and easy to look after, with a short life cycle. S. pombe is a particularly useful model organism because:
- It is a single cell with just 5000 genes (compared to 20 000 in human cells);
- The genome is easy to manipulate – genes can be added, mutated and removed. They can also be tagged with fluorescent markers, enabling proteins to be visualised using specialised microscopes;
- The genome is usually haploid (there is only one copy of each chromosome), which means that genetic changes will be expressed in the phenotype and not masked by another gene;
- The cell cycle is simple and it is easy to tell what stage the cell is in by simply looking at it;
- Many of the genes in S. pombe have homologues (equivalents) in human cells;
- Cells are quick, cheap and easy to grow and can be stored at –80 °C for many years.
Unfortunately, human cells are not ideal for cell-cycle research. The human genome contains about 20 000 genes, many of which have more than one function. In addition, human cells and DNA can be difficult to manipulate in the lab, making it tricky to unpick the function of one particular gene in this way. The yeast S. pombe has provided a solution. This single-celled organism, also known as fission yeast, is just 4 µm wide and 14 µm long and only has about 5000 genes.
S. pombe was first described in 1893 by German scientist Paul Lindner, who discovered the yeast in East African millet beer. Because of this, he named it after the Swahili word for beer, pombe. Researchers began studying the genetics and cell cycle of S. pombe in the 1950s. It is ideal for research for many reasons: it is easy to grow and non-pathogenic, and the cells are big enough to see under a microscope, contain just three chromosomes and divide every 2–4 hours (see box).
A major strength of S. pombe as a model organism is the ease with which its genome can be manipulated – specific genes can be completely removed or extra DNA can be added.
Scientists can grow these genetically altered cells and the resulting phenotype (the organism’s observable characteristics) then helps them to understand the function of the altered gene. S. pombe cells are also haploid, meaning there is only one copy of each gene. This makes it even easier to look at gene function. In a diploid cell (where there are two copies of each gene), great care needs to be taken to ensure both copies of a gene are changed in order for the phenotype to be meaningful. Having just one copy of a gene, as in S. pombe, makes this process simple.
In addition, S. pombe has a well characterized cell cycle and its regular rod shape makes it ideal for studies of growth and division. The cells maintain their rod shape and elongate at the ends until a precise cell length has been reached. The DNA is replicated and one copy of the genome moves to each half of the cell, then a septum (brightly stained with a fluorescent dye) forms across the centre of the cell and it divides (see figure 1). This cycle repeats, with each cell growing and dividing into two daughter cells as long as the conditions are suitable for growth (e.g. plentiful nutrients are present).
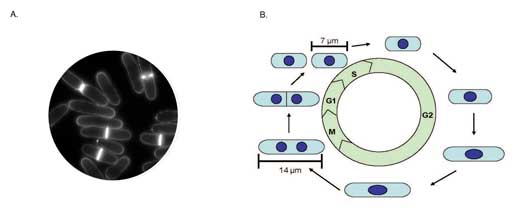
Image courtesy of Louise Weston
In the 1970s, Paul Nurse and his colleagues began isolating and characterising cell-cycle mutants in S. pombe – work that ultimately led to a Nobel Prize in 2001. Initially the researchers looked for genetic alterations that caused cell death or unusually elongated cells that failed to divide (Nurse et al., 1976). Figure 2(b) shows elongated cells that have failed to divide following inhibition of a gene called cdc (which stands for cell-division cycle).
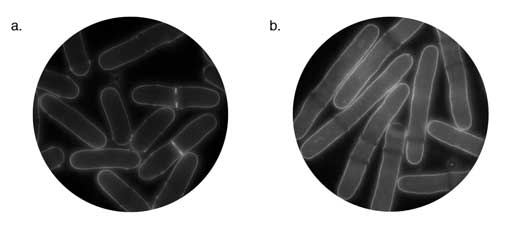
Image courtesy of Louise Weston
The benefits of model organisms become clear when the research advances our understanding of similar processes in humans. Many of the genes found in S. pombe to regulate DNA damage and repair, checkpoint controls and the cell cycle have homologues (equivalents) in human cells. For example, after identifying the gene cdc2 in S. pombe, Nurse and his colleagues went on to find the corresponding gene, cdk1 (cyclin dependent kinase 1), in humans (Lee & Nurse, 1987). This gene codes for a protein that initiates and regulates cell division in humans. Mutations in this gene can trigger unscheduled cell division and the duplication or deletion of chromosome sections, which may lead to cancer. Today, the therapeutic benefit of selective CDK inhibition in human cancer treatment is being studied. The opportunities go beyond addressing cancer – in fact, homologues of at least 50 S. pombe genes are associated with human diseases as varied as cystic fibrosis, hereditary deafness and type 2 diabetes.
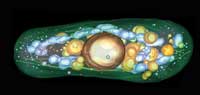
inside of fission yeast. There
is still more to understand
about the inside of this
fungus.
Image courtesy of Johanna
Höög, EMBL
S. pombe is so essential for research that, in 2002, it was the sixth eukaryotic organism to have its entire genome sequence published. This has led to the development of more important tools for studying S. pombe. For example, a library of 3308 strains of S. pombe has been created, each of which is missing one of the yeast’s non-essential genes. In addition, each protein in S. pombe has been tagged with green fluorescent protein (which glows bright green under ultraviolet light), allowing its exact location within the cell to be observed with a fluorescent microscope. These tools will make S. pombe an even more useful model organism for investigating gene function (Yanagida, 2002).
We still don’t fully understand how processes such as growth and cell division are controlled, and global controls over cell growth are particularly poorly understood. There is much we can learn from this simple single-celled organism about the complex processes that are so important to our healthy development as well as the development of diseases like cancer.
References
- Lee MG, Nurse P (1987) Complementation used to clone a human homologue of the fission yeast cell cycle control gene cdc2. Nature 327: 31-35
- Nurse P, Thuriaux P, Nasmyth K (1976) Genetic control of the cell division cycle in the fission yeast Schizosaccharomyces pombe. Molecular & General Genetics 146: 167-178
- Yanagida M (2002) The model unicellular eukaryote, Schizosaccharomyces pombe. Genome Biology 3: comment2003.4
Resources
- For more information about S. pombe and how it can be used in scientific research, visit PombeNet, a resource produced by the Forsburg Lab in California.
- The Cancer Research UK website offers accessible information on all the major cancers and current research.
- For more information on how genetic mutations cause diseases, see:
- Patterson L (2009) Getting a grip on genetic diseases. Science in School 13: 53-58.
- For a classroom activity to discuss the ethics of knowing what your genes have in store for you, including the possibility of cancer, see:
- Meet the Gene Machine: stimulating bioethical discussions at school. Science in School 9: 34-38.
Review
Yeasts and fungi are with us, in us, and all around us. Students should learn about them while at school. Textbooks recommend safe handling of fungi growth but often provide no further detailed experimental procedures to follow. Starting with the usage of fungi in everyday food conservation and production will ignite students’ interest in the topic. (Teachers should shy away from any practical using mould experiments, even under enclosed petri dishes.)
Using a safe yeast strain (e.g. baking with yeast, fermenting food or brewing beer or wine) in a well-defined experiment should encourage any biology teacher to give students the opportunity not only to learn good lab habits but also to understand why certain procedures need to be followed precisely and recorded correctly before one can draw any conclusions. This is how scientific practice should be taught before the students are introduced to the article, which is exemplary for a scientific report. The facts presented can lead to virtually planning experiments by using the described model organism. The questions concluding the report will generate more hypotheses for further research and discussions in class.
Friedlinde Krotscheck, Germany





