Supporting materials
Download
Download this article as a PDF

Russ Hodge from the European Molecular Biology Laboratory in Heidelberg, Germany, reports on the first complete survey of 'molecular machines' in yeast.

In 1901, Franz Hofmeister compared the cell to a factory, able to take in raw elements and convert them into the necessities of life; he even suggested that the sub-compartments of cells that had been identified under the microscope might be responsible for specific types of conversions.
The factory analogy has persisted through a century of discoveries about the functions of molecules. Proteins were described as ‘worker molecules’ and chemical processes as ‘assembly lines’. Unlike a car factory, however, where machines usually remain bolted to the floor and are only changed as new models come into fashion, the cell continually retools itself. Proteins are simultaneously workers and components of intricate robots that are continually assembled and taken apart; often, the same molecule can be found in several machines.
The full extent of this flexible organisation has only become clear through a recent study published in the journal Nature (Gavin et al., 2006). Previously, scientists had a very limited view of the machines and their components. “The situation was like coming into a factory and finding parts of single machines scattered across the floor,” says lead researcher Anne-Claude Gavin, a scientist at the European Molecular Biology Laboratory (EMBL) in Heidelberg, Germany. “We have known what some machines do, and a bit about how they operate, but there was really no view of the whole context.”

Scientists had already started to puzzle together the construction of yeast machines based on single pieces, using a method called a two-hybrid screen. This matches every yeast protein to every other, like completely disassembling everything in a car factory and trying to fit pieces together one-by-one. The method has generated a wealth of useful information but also many false leads.
It might be physically possible to insert a gear stick into an exhaust pipe, but that doesn’t mean it ever happens in a functioning automobile. With 6500 parts to deal with – approximately the number of proteins encoded in the yeast genome – the one-by-one method provides a very limited view of complete machines, let alone the complete factory.
An alternative would be to start with complete machines and then analyse the molecules that compose them. But methods of extracting proteins from cells usually break complexes apart. Then, several years ago, Bertrand Séraphin’s lab at EMBL invented a process called tandem affinity purification (TAP), a method which fishes single molecules from cells along with any machines attached to them, intact. The components of complexes could then be analysed using mass spectrometry – a method that fragments proteins and ‘weighs’ the pieces. Since each protein has a unique composition, mass spectrometry gives scientists measurements that can be matched by computer to the profile of a specific molecule. Working with scientists at EMBL, the company Cellzome decided to tackle the entire yeast genome using these methods. Thousands of experiments later, this has produced the first full scan of the genes of a eukaryotic cell, searching for molecular machines.
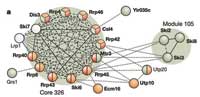
The study revealed 491 complexes, 257 of them wholly new. The rest were familiar from other research, but now virtually all of them were found to have new components.Is the list exhaustive? “We estimate there may about 300 more,” Anne-Claude says. “Some complexes may appear only when particular conditions are used to grow the yeast, and others may not be discoverable with this method of extracting them.” For example, it has been notoriously difficult to purify complexes attached to the cell membrane. The researchers adapted the method for doing so, and found 74 new complexes involving membrane proteins, but Anne-Claude is sure that many more exist.
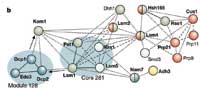
A parts list is only the beginning: the scientists also want to know where the complexes are stationed in the cell, what they do, and how they function. Sometimes these questions can be answered from the components alone. A complex with three proteins that respond to heat undoubtedly plays a role in helping the organism adapt to changes in temperature. Other complexes could be linked to processes such as binding to DNA, or assisting in the refinement of other molecules.
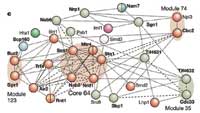
The information has also provided new insights into how the cell manages the incredibly complicated task of putting complexes together, and this says something important about the biology of yeast and other organisms. “Does a cell pre-assemble machines and have them on hand, or are they built from scratch when something happens?” Anne-Claude says. “In other words, how are the machines – and the factory as a whole – really managed? We didn’t know, but now we can say a lot about this. To do that, we had to find new ways of understanding the data.”
Road atlases often contain a chart showing the driving distances between cities. For example, 1150 kilometres separate Rome and Heidelberg; Heidelberg to Cambridge is 2045 kilometres (counting a transit by ferry). From such a chart and a bit more information, you could sort cities into regions, states, and countries. To understand the inner workings of protein complexes, says Patrick Aloy, scientists would like to have such a map of molecules.
Patrick was a member of Rob Russell’s group at EMBL, where computational techniques are used to understand the inner workings of protein complexes. Combining information on protein shapes and functions with data on how they bind to other molecules has permitted the scientists to start drawing ‘technical diagrams’ of machines. But sketchy knowledge has restricted the scope of these efforts.
“Imagine replacing a chart of distances with a table that tells you only yes or no – can you get there from here?” Patrick says. “From that information you wouldn’t be able to draw a very meaningful map, but that type of study is what we’ve had so far. Well, now we’ve produced something more like the distance chart – each pair of proteins has a value which gives the likelihood that they are found together in purifications.”
That information has now been converted into a map of the factory floor, complete with finished and partial machines, prefabricated parts, and snap-on modules. “What you discover is that most complexes have a core set of components that are almost always found together and others that come and go,” Rob says. “You can think of the cores as crucial, prefabricated parts of machines that are kept on hand, with temporary modules added on as the need arises.”
The function of such modules, Anne-Claude says, may be to alter the job of the core machine, to link it to other things going on in the factory, to switch it on or shut it down. “This has several very important effects. First, it gives the cell a way to carry out a large number of tasks with a limited number of basic machines. That gives it quite a bit of flexibility. Secondly, it means that in an emergency, the cell doesn’t have to build all the machines it needs from scratch. It only has to produce a few really essential parts. The other side of that is that it may be relatively simple for the cell to control a quite sophisticated machine, just by supplying or blocking the delivery of a critical piece.”
There is an important connection to evolution because generally, certain types of machines and their basic components have been conserved over hundreds of millions of years as new species arose. Peer Bork’s group at EMBL has helped in investigating this question.
“If you compare what goes on in yeast and our own cells, you find many of the same machines, using the same basic components to do the same things,” Peer says. “The complexes reflect how evolution works – as variations on a theme. You don’t find every species inventing a new way of doing things; instead, they’re refining what they’re up to by adding on specialised modules, or slightly changing the way the whole thing is regulated.”
The study reveals a great deal about individual machines and how they work together. Yet much remains to be learned about their work in living cells – where many of them are located, and how many copies of each machine are at work at any given time. Structural information about the complexes is helping to answer some of these questions, because it gives scientists an idea about the overall shape of a complex. This means it can be looked for under the microscope.
“Even with electron microscopy, protein complexes are fuzzy spots that are difficult or impossible to identify,” Anne-Claude says. “But with good shape information, we might be able to put names to some of the shapes.”
Health reflects the operation of the entire cell in the context of the organism. The level of molecular machines is crucial in influencing that and keeping things in balance. What the scientists have accomplished is changing what we think about not only how individual machines work and are regulated, but how they function together. That will obviously be crucial as scientists try to guide organisms from a diseased state to a healthy one.
In the field of life sciences, the word proteome has become very popular, like other words ending with “-ome”, but it is rare to find it explained as clearly as it is in this article.
Beginning with the metaphor of the cell as a factory, Russ Hodge leads the reader into its complex protein machinery using a plain and captivating style, with vivid examples from everyday life.
Another engaging feature of the article is the description of methods and strategies used by researchers to investigate the structure, function and evolution of those incredibly tiny objects that are cell protein complexes.
I recommend this article to teachers and students of upper secondary school within the curriculum of biochemistry and/or cell biology. In fact, it can be used for classroom teaching or for autonomous investigation of the subject by students.
Finally I would stress that this article, in spite of the simple and friendly style, offers an illuminating and synthetic view of a very complex issue: with the exciting taste of the adventure of discovery, it gives an excellent example of high-level science popularisation, which is especially needed at present to encourage scientific vocations.
Giulia Realdon, Italy
Download this article as a PDF