How do muscles produce work? Using optical tweezers to study molecular machines Understand article
Alexandre Lewalle from King’s College, London, UK, pushes back the frontiers of our knowledge of motors – at the molecular level.
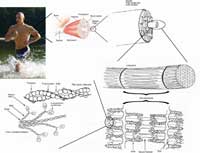
actin filaments are the most
basic constituents of muscle
tissue. They form
overlapping filaments in the
cells that slide past one
another to make the muscle
contract.
Image courtesy of Raul654;
image source: Wikimedia
Commons
There is little mystery in how we lift a shopping bag: we bend, grasp and pull. As we know, our body transforms chemical energy into mechanical work, like a motor powered by fuel. But surely our body isn’t an engine made of gears and pistons – or is it?
As it turns out, at the microscopic level, every cell in our muscles is made up of millions of proteins that indeed are tiny motors comprising only one type of molecule, no bigger than a few nanometres. These proteins, called myosin, are arranged in a regular pattern and work in unison to produce forces and movement many times greater than those they achieve individually.
There is currently much enthusiasm for nanotechnology, the field that explores and seeks to control the microscopic world. In my lab at King’s College, London, we use an apparatus called ‘optical tweezers’ to manipulate and study these motors individually. Microscopic man-made machines may still be the stuff of science fiction, but through experiments of this kind, nature may well teach us our first lesson.
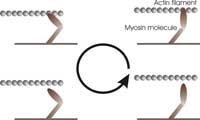
mechanical work by pulling
on the actin filament in a
cyclical manner: (1) binding
to the filament, (2) pulling on
the filament, (3) releasing the
filament, and (4) refuelling
(one molecule of ATP).
Image courtesy of Alexandre
Lewalle
By understanding how these molecules work in detail, we can gain insight into the ways that nature, through evolution, has engineered many different forms of life from remarkably few templates. Many kinds of single-molecule ‘machines’ exist in nature, and the tasks they perform are as varied as they are vital. They enable living cells to function, move and reproduce. For example, some of them transport nutrients across the cell by ‘walking’ along an intracellular railroad network. Molecular-sized motors probably play a crucial role in cell division by separating chromosomes after they have replicated. Other types of motors rotate the flagellum that propels sperm cells and some bacteria. Many of these motors are structurally very similar, and yet they perform distinct tasks.
Muscle contraction is achieved by sliding two kinds of filaments past one another. One of these filaments is made of myosin (the motor), the other of a substance called actin (see figure above). The two kinds of filaments overlap in the muscle cells to maximise their interactions.
Just like a more familiar engine, myosin operates in a cyclical manner, as shown in the figure (right), to ensure energy is used efficiently. During each cycle, the myosin molecules first bind to the actin filament, bridging the gap between the two filaments. Then, they ‘burn’ fuel by reacting with one ATP molecule (the fuel of most biological processes). This reaction releases energy and makes the myosin molecules undergo a transformation that causes them to swing their ‘lever arm’ structure and pull on the actin filament. They then release the actin filament and swing the lever arm back to its original position, ready for the next cycle.
During each cycle, the two filaments slide past each other by a short distance – this is the basis of muscle contraction. The forces and movements produced by a single cycle are minute, but the combination of millions of myosin molecules acting simultaneously amplifies the effect by many orders of magnitude.
Biologists have been studying muscle and myosin for many decades, but for all our present knowledge, there still remains a vast region of mystery: how strongly do the individual myosin molecules pull and over what distance? How is the duration of one cycle affected by the chemical environment? We know that the total work done by the myosin molecule is strongly affected by the external force against which the motor has to pull, but the relationship between them is still poorly understood. Because of their small size, motor molecules like myosin react to their environment very differently than would larger molecules. For example, in contrast to a car engine, the effects of viscosity in the environment and of the continual bombardment of water molecules (the phenomenon known as Brownian motion) are considerable. At the molecular scale, moving through water actually feels more like what we would experience when swimming in turbulent honey!
Scientists are insatiably curious. Theorists devise models to suggest how muscles work, but at the end of the day, only experiments will really push our understanding further. However, a detailed understanding of myosin cannot be gained by looking at whole muscles because what we see is only the overall effect of many myosin motors acting independently. What is needed instead is a way to control and examine the myosin molecules in isolation. But this is a monstrous challenge in itself. These are obviously not objects you can simply hold with a steady hand!
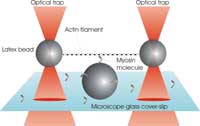
A single filament of actin is
stretched between two
plastic beads held in optical
tweezers. The myosin
molecule can then bind to the
filament, and its working
stroke is detectable from the
positions of the two trapped
beads.
Image courtesy of Alexandre
Lewalle
It is only in recent years that scientists have developed tools that allow them to manipulate single molecules, opening up new avenues of research and blurring the boundaries of traditional disciplines like physics and biology. The ‘optical tweezers’, or ‘optical trap’, use the effect by which a laser beam, when focused by the strong objective lens of a microscope, traps microscopic objects. This trapping effect works by exploiting the fact that photons carry momentum and therefore exert minute forces on particles. A small displacement of a particle away from the focus causes a deflection of the beam, and hence, by Newton’s Third Law, the particle experiences a restoring force back towards the trap centre. This sensitive technique provides a direct handle on the individual molecules. The appeal of applying it to study myosin is that the forces exerted by the laser are of similar magnitude to those exerted by myosin itself.
Our experiment at King’s College is designed to recreate, under the microscope, the basic unit of a contractible muscle. The aim is to investigate how a single myosin molecule responds to an external force applied to an actin filament via the optical tweezers. First we produce two traps by shining two laser beams into our microscope objective (see figure). Each trap is then made to catch one micron-sized plastic bead that is specially coated to bind actin filaments. A single filament is caught and stretched between the beads, producing a kind of dumbbell. We now steer this dumbbell towards another bead that is stuck to the surface of our microscope cover slip and coated with myosin molecules. When the conditions are just right, we manage to make just one myosin molecule, located near the top of the central bead, interact with the actin filament in the dumbbell, just as it would do in a whole muscle fibre.

apparatus.
Image courtesy of Alexandre
Lewalle
Interactions between myosin and actin can be detected by monitoring the positions of the two beads in the dumbbell. When myosin binds and pulls on the myosin (as illustrated in the figure above), the beads are displaced from the trap centres by a small but measurable distance. This displacement tells us not only how far the myosin motor has pulled the actin, but also how much force it has exerted against the strength of the two traps. Our measurements suggest that the average displacement from a single myosin molecule is about 10 nanometres (a millionth of a centimetre), with typical forces of only a few piconewtons (10-12 newtons). These values are important parameters in any theoretical model that tries to explain the physiology of muscles from the bottom upwards.
It goes without saying that these experiments require a good dose of patience. They also require the collaboration of scientists with very different domains of expertise. Biologists are crucial to connect the results of the experiments to the known physiological behaviour of muscles, and to ensure that the project remains on the right track. Physicists are equally important in properly setting up the optical and electronic components of the apparatus and in analysing the results quantitatively. As in any area of research, the frontiers of our knowledge are only extended by trying out new things and by being endlessly curious and inquisitive. So the next time you think there is nothing simpler than throwing a basketball, think again.
Review
Nanotechnology is a fashionable subject in science fiction as well as in cutting-edge research and, in this interesting article, Alexandre Lewalle tells us about nanomotors in muscle cells (the tiny actin and myosin filaments) and about the ways in which they are investigated by means of nanotools (the optical tweezers).
The article, written in an enjoyable style with clear examples and vivid metaphors, is suitable for biology teachers willing to update their knowledge and for upper secondary school students interested in the forefront of research.
While the addressed topic is not part of the usual biology curricula, it is related to physiology and biochemistry, and it is a good example of current research in the biomedical field. It can be used to foster interest in science and encourage young people to continue their scientific studies.
Giulia Realdon, Italy





